|
|
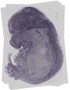
Ttc22 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
|
EMAGE:11187
view entry |
wild-type |
- |
ISH |
 |
|
|
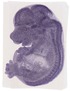
Pcdhb3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.341 |
| Go to annotation details |
 |
 |
facial VII ganglion |
 |
hindbrain |
 |
trigeminal V nerve |
|
 |
forebrain |
 |
neural retina |
 |
trigeminal V ganglion |
 |
glossopharyngeal IX ganglion |
 |
vestibulocochlear VIII ganglion |
 |
spinal cord |
 |
midbrain |
 |
dorsal root ganglion |
|
|
EMAGE:12162
view entry |
wild-type |
- |
ISH |
 |
|
|
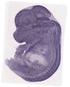
Ablim3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.341 |
| Go to annotation details |
 |
 |
facial VII ganglion |
 |
dorsal root ganglion |
 |
heart valve |
|
 |
vestibulocochlear VIII ganglion |
 |
diencephalon |
 |
glossopharyngeal IX ganglion |
 |
vagus X ganglion |
 |
neural retina |
 |
telencephalon |
 |
trigeminal V nerve |
 |
midbrain |
 |
hindbrain |
 |
spinal cord |
 |
trigeminal V ganglion |
|
|
EMAGE:12076
view entry |
wild-type |
- |
ISH |
 |
|
|
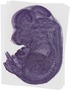
Acat2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.339 |
| Go to annotation details |
 |
 |
vestibulocochlear VIII ganglion |
 |
thoracic ganglion |
 |
chondrocranium |
|
 |
liver lobe |
 |
vagus X ganglion |
 |
glossopharyngeal IX ganglion |
 |
stomach |
 |
cervico-thoracic ganglion |
 |
head mesenchyme |
 |
upper jaw incisor |
 |
dorsal root ganglion |
 |
hindgut |
 |
lower jaw incisor |
 |
midgut |
 |
foregut-midgut junction |
 |
maturing glomerular tuft |
 |
trigeminal V ganglion |
 |
cervical ganglion |
 |
olfactory cortex marginal layer |
 |
vibrissa |
 |
facial VII ganglion |
|
|
EMAGE:11188
view entry |
wild-type |
- |
ISH |
 |
|
|
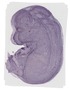
Mettl7b |
 |
 |
 |
TS23 |
14.5 dpc |
0.338 |
- |
EMAGE:11190
view entry |
wild-type |
- |
ISH |
 |
|
|
Tceal8 |
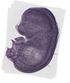 |
 |
 |
TS23 |
14.5 dpc |
0.338 |
- |
EMAGE:19410
view entry |
wild-type |
- |
ISH |
 |
|
|
Ankrd1 |
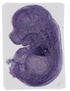 |
 |
 |
TS23 |
14.5 dpc |
0.337 |
| Go to annotation details |
 |
 |
heart left ventricle |
 |
heart atrium |
 |
vertebral axis musculature |
|
 |
heart right ventricle |
 |
tongue muscle |
|
|
EMAGE:11742
view entry |
wild-type |
- |
ISH |
 |
|
|
Ywhaq |
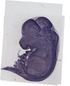 |
 |
 |
TS23 |
14.5 dpc |
0.337 |
- |
EMAGE:9163
view entry |
wild-type |
- |
ISH |
 |
|
|
Mtmr1 |
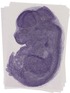 |
 |
 |
TS23 |
14.5 dpc |
0.337 |
|
EMAGE:12313
view entry |
wild-type |
- |
ISH |
 |
|
|
Rfx5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.337 |
- |
EMAGE:16589
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
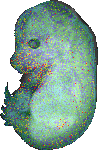 download query region as wlz file
download query region as wlz file