|
|
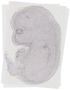
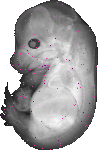
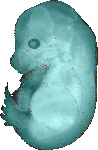
Kcnc3 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
|
EMAGE:12004
view entry |
wild-type |
- |
ISH |
 |
|
|
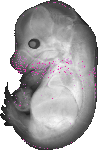
Moxd2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.026 |
- |
EMAGE:15623
view entry |
wild-type |
- |
ISH |
 |
|
|
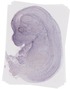
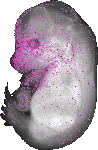
Amdhd2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.023 |
|
EMAGE:11242
view entry |
wild-type |
- |
ISH |
 |
|
|
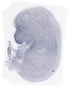
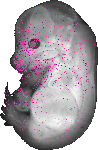
Gmppa |
 |
 |
 |
TS23 |
14.5 dpc |
0.022 |
- |
EMAGE:10900
view entry |
wild-type |
- |
ISH |
 |
|
|
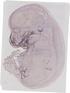
Col17a1 |
 |
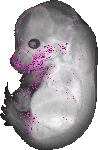 |
 |
TS23 |
14.5 dpc |
0.021 |
| Go to annotation details |
 |
 |
vibrissa |
 |
epidermis |
 |
cornea epithelium |
|
 |
pharyngo-tympanic tube |
 |
oral epithelium |
 |
larynx |
 |
esophagus |
 |
nasal cavity respiratory epithelium |
 |
naris |
|
|
EMAGE:10011
view entry |
wild-type |
- |
ISH |
 |
|
|
Mir204 |
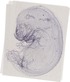 |
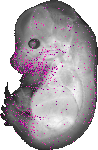 |
 |
TS23 |
14.5 dpc |
0.021 |
| Go to annotation details |
 |
 |
pharynx |
 |
pharyngo-tympanic tube |
 |
stomach |
|
 |
esophagus |
 |
vibrissa |
 |
epidermis |
 |
lateral ventricle choroid plexus |
 |
nasal cavity olfactory epithelium |
 |
submandibular gland primordium |
 |
urethra of male |
 |
larynx |
 |
oral epithelium |
 |
metencephalon part of 4th ventricle choroid plexus |
 |
cornea epithelium |
|
|
EMAGE:13777
view entry |
wild-type |
- |
ISH |
 |
|
|
A530013C23Rik |
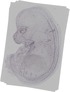 |
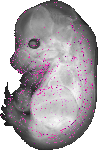 |
 |
TS23 |
14.5 dpc |
0.021 |
| Go to annotation details |
 |
 |
rib |
 |
femur |
 |
pelvic girdle skeleton |
|
 |
scapula |
 |
fibula |
 |
radius |
 |
tibia |
 |
humerus |
 |
Meckel's cartilage |
 |
axial skeleton |
|
|
EMAGE:16004
view entry |
wild-type |
- |
ISH |
 |
|
|
3830406C13Rik |
 |
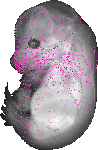 |
 |
TS23 |
14.5 dpc |
0.021 |
- |
EMAGE:14677
view entry |
wild-type |
- |
ISH |
 |
|
|
Rhbdf1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.021 |
- |
EMAGE:10876
view entry |
wild-type |
- |
ISH |
 |
|
|
Jarid2 |
 |
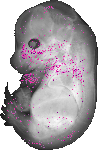 |
 |
TS23 |
14.5 dpc |
0.021 |
- |
EMAGE:14540
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file