|
|
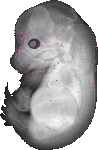
Otub2 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:14579
view entry |
wild-type |
- |
ISH |
 |
|
|
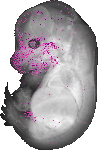
Alx4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.040 |
| Go to annotation details |
 |
 |
telencephalon meninges |
 |
extrinsic ocular muscle |
 |
submandibular gland primordium |
|
 |
eye |
 |
mandible |
 |
diencephalon roof plate |
 |
genital tubercle of male |
 |
vibrissa |
 |
viscerocranium |
|
|
EMAGE:20057
view entry |
wild-type |
- |
ISH |
 |
|
|
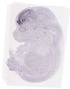
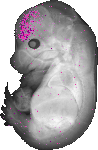
Eif1b |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
|
EMAGE:12794
view entry |
wild-type |
- |
ISH |
 |
|
|
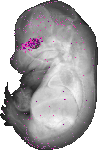
Mybl1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.034 |
| Go to annotation details |
 |
 |
medulla oblongata basal plate ventricular layer |
 |
adrenal gland |
 |
neural retina |
|
 |
testis |
 |
vomeronasal organ |
 |
pons ventricular layer |
 |
midbrain ventricular layer |
 |
nasal cavity olfactory epithelium |
|
|
EMAGE:13810
view entry |
wild-type |
- |
ISH |
 |
|
|
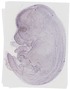
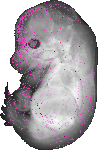
Marveld2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.034 |
| Go to annotation details |
 |
 |
midgut |
 |
nasal capsule |
 |
bladder |
|
 |
pancreas |
 |
hindgut |
 |
kidney pelvis |
 |
foregut-midgut junction |
 |
stomach |
 |
kidney calyx |
|
|
EMAGE:11045
view entry |
wild-type |
- |
ISH |
 |
|
|
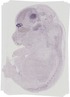
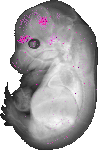
Kctd6 |
 |
 |
 |
TS23 |
14.5 dpc |
0.034 |
| Go to annotation details |
 |
 |
vibrissa |
 |
dorsal root ganglion |
 |
trigeminal V ganglion |
|
 |
olfactory cortex marginal layer |
 |
thalamus mantle layer |
|
|
EMAGE:11198
view entry |
wild-type |
- |
ISH |
 |
|
|
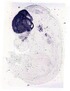
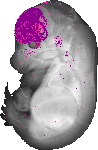
Foxg1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
- |
EMAGE:9057
view entry |
wild-type |
- |
ISH |
 |
|
|
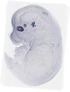
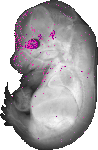
Rax |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
|
EMAGE:10998
view entry |
wild-type |
- |
ISH |
 |
|
|
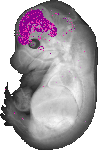
Eomes |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
- |
EMAGE:9323
view entry |
wild-type |
- |
ISH |
 |
|
|
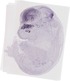
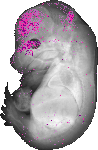
Insm1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
| Go to annotation details |
 |
 |
telencephalon ventricular layer |
 |
vomeronasal organ |
 |
diencephalon lateral wall ventricular layer |
|
 |
metencephalon rest of alar plate marginal layer |
 |
neural retina |
 |
midbrain ventricular layer |
 |
adrenal gland |
 |
nasal cavity olfactory epithelium |
 |
olfactory cortex ventricular layer |
 |
pancreas |
 |
diencephalon lateral wall mantle layer |
 |
hypothalamus ventricular layer |
|
|
EMAGE:21644
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file