|
|
Pan2 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:17681
view entry |
wild-type |
- |
ISH |
 |
|
|
Plod2 |
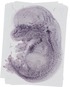 |
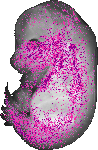 |
 |
TS23 |
14.5 dpc |
0.038 |
| Go to annotation details |
 |
 |
otic capsule |
 |
axial skeleton |
 |
axial skeleton tail region |
|
 |
sternum |
 |
foot mesenchyme |
 |
trachea |
 |
nasal septum |
 |
tongue |
 |
rib |
 |
temporal bone petrous part |
 |
clavicle |
 |
basisphenoid bone |
 |
exoccipital bone |
 |
viscerocranium |
|
|
EMAGE:16891
view entry |
wild-type |
- |
ISH |
 |
|
|
Lrig3 |
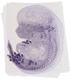 |
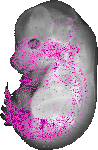 |
 |
TS23 |
14.5 dpc |
0.037 |
- |
EMAGE:13604
view entry |
wild-type |
- |
ISH |
 |
|
|
Slit3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
- |
EMAGE:13229
view entry |
wild-type |
- |
ISH |
 |
|
|
Parva |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
- |
EMAGE:9664
view entry |
wild-type |
- |
ISH |
 |
|
|
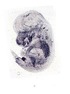
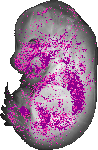
Bambi |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
- |
EMAGE:9050
view entry |
wild-type |
- |
ISH |
 |
|
|
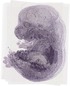
Csrp1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
| Go to annotation details |
 |
 |
lower jaw molar |
 |
submandibular gland primordium |
 |
stomach |
|
 |
palatal shelf |
 |
axial skeleton |
 |
ureter |
 |
metanephros |
 |
esophagus |
 |
testis |
 |
bladder |
 |
hindgut |
 |
midgut |
 |
vibrissa |
 |
clavicle |
 |
axial skeleton tail region |
 |
upper jaw incisor |
 |
lung |
 |
trachea |
 |
aorta |
 |
upper jaw molar |
 |
tail mesenchyme |
 |
lower jaw incisor |
 |
tongue muscle |
|
|
EMAGE:12991
view entry |
wild-type |
- |
ISH |
 |
|
|
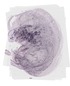
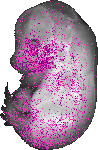
Ptgfr |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
| Go to annotation details |
 |
 |
Meckel's cartilage |
 |
upper jaw incisor |
 |
lower jaw molar |
|
 |
nose |
 |
upper jaw molar |
 |
orbito-sphenoid |
 |
lower jaw incisor |
|
|
EMAGE:21275
view entry |
wild-type |
- |
ISH |
 |
|
|
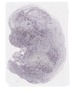
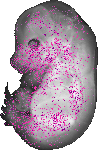
Tgif1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
| Go to annotation details |
 |
|
|
 |
renal cortex |
 |
diencephalon lateral wall ventricular layer |
 |
bladder |
 |
telencephalon ventricular layer |
 |
vibrissa |
 |
esophagus |
 |
lung |
 |
urethra of male |
 |
rectum |
 |
midbrain ventricular layer |
|
|
EMAGE:13961
view entry |
wild-type |
- |
ISH |
 |
|
|
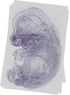
Pdgfa |
 |
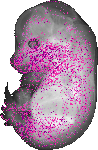 |
 |
TS23 |
14.5 dpc |
0.036 |
| Go to annotation details |
 |
 |
upper jaw molar |
 |
vomeronasal organ |
 |
nasal cavity olfactory epithelium |
|
 |
tail paraxial mesenchyme |
 |
pharyngo-tympanic tube |
 |
lower jaw molar |
 |
vibrissa |
 |
forearm rest of mesenchyme |
 |
trachea |
 |
thymus primordium |
 |
right lung |
 |
upper jaw incisor |
 |
lower leg rest of mesenchyme |
 |
lower jaw incisor |
 |
metencephalon part of 4th ventricle choroid plexus |
 |
external naris |
 |
naso-lacrimal duct |
 |
vertebral axis musculature |
 |
submandibular gland primordium |
 |
upper arm muscle |
 |
hand |
 |
midgut |
 |
orbito-sphenoid |
 |
axial skeleton tail region |
 |
choroid invagination |
 |
pharynx epithelium |
 |
diencephalon roof plate |
 |
tongue epithelium |
 |
kidney calyx |
 |
upper leg muscle |
 |
seminiferous cord |
 |
stomach |
 |
conjunctival sac |
 |
diaphragm |
 |
epidermis |
 |
posterior naris |
 |
tongue muscle |
 |
eye skeletal muscle |
 |
left lung |
 |
oral epithelium |
 |
urethra of male |
 |
esophagus epithelium |
 |
anterior naris |
 |
foot |
|
|
EMAGE:14220
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
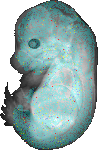 download query region as wlz file
download query region as wlz file