|
|
Maged1 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
mesenchyme |
 |
urinary system |
 |
sensory organ system |
|
 |
integumental system |
 |
body cavity or lining |
 |
alimentary system |
 |
vertebral axis musculature |
 |
respiratory system |
 |
reproductive system |
 |
cardiovascular system |
 |
gland |
 |
tail |
 |
nervous system |
 |
limb |
|
|
EMAGE:20278
view entry |
wild-type |
- |
ISH |
 |
|
|
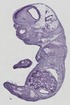
Ctsb |
 |
 |
 |
TS23 |
14.5 dpc |
0.886 |
- |
EMAGE:11101
view entry |
wild-type |
- |
ISH |
 |
|
|
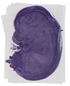
Iqcc |
 |
 |
 |
TS23 |
14.5 dpc |
0.883 |
- |
EMAGE:11158
view entry |
wild-type |
- |
ISH |
 |
|
|
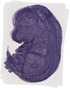
Plk3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.882 |
- |
EMAGE:11829
view entry |
wild-type |
- |
ISH |
 |
|
|
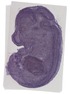
Pcyt2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.881 |
|
EMAGE:11498
view entry |
wild-type |
- |
ISH |
 |
|
|
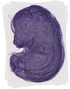
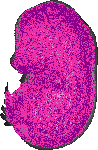
Ppp1r11 |
 |
 |
 |
TS23 |
14.5 dpc |
0.880 |
|
EMAGE:12326
view entry |
wild-type |
- |
ISH |
 |
|
|
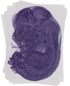
Noc2l |
 |
 |
 |
TS23 |
14.5 dpc |
0.880 |
- |
EMAGE:11163
view entry |
wild-type |
- |
ISH |
 |
|
|
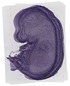
Gbp8 |
 |
 |
 |
TS23 |
14.5 dpc |
0.880 |
|
EMAGE:12381
view entry |
wild-type |
- |
ISH |
 |
|
|
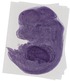
Ubac1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.880 |
| Go to annotation details |
 |
 |
liver right lobe |
 |
vibrissa |
 |
glossopharyngeal IX ganglion |
|
 |
dorsal root ganglion |
 |
liver left lobe |
 |
facial VII ganglion |
 |
vestibulocochlear VIII ganglion |
 |
trigeminal V ganglion |
|
|
EMAGE:11102
view entry |
wild-type |
- |
ISH |
 |
|
|
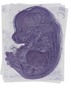
Fam55b |
 |
 |
 |
TS23 |
14.5 dpc |
0.880 |
| Go to annotation details |
 |
 |
heart ventricle |
 |
stomach |
 |
cranial muscle |
|
 |
eye skeletal muscle |
 |
otic capsule |
 |
nucleus pulposus |
 |
arm |
 |
spinal ganglion |
 |
neurohypophysis |
 |
labyrinth |
 |
hip |
 |
viscerocranium |
 |
pelvic girdle muscle |
 |
forelimb |
 |
pituitary gland |
 |
trigeminal V ganglion |
 |
urethra of male |
 |
shoulder mesenchyme |
 |
upper leg |
 |
heart |
 |
leg |
 |
axial muscle |
 |
shoulder |
 |
liver |
 |
lung |
 |
gut |
 |
pectoral girdle and thoracic body wall muscle |
 |
naso-lacrimal duct |
 |
dorsal root ganglion |
 |
vibrissa |
|
|
EMAGE:10116
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
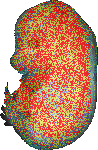 download query region as wlz file
download query region as wlz file