|
|
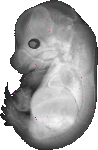
Rnasel |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:8458
view entry |
wild-type |
- |
ISH |
 |
|
|
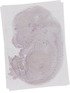
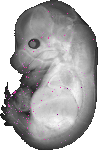
Ube2a |
 |
 |
 |
TS23 |
14.5 dpc |
0.014 |
- |
EMAGE:9917
view entry |
wild-type |
- |
ISH |
 |
|
|
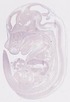
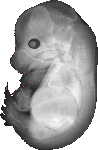
Evi2a |
 |
 |
 |
TS23 |
14.5 dpc |
0.012 |
- |
EMAGE:9617
view entry |
wild-type |
- |
ISH |
 |
|
|
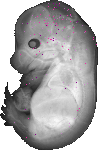
Slc41a1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.011 |
| Go to annotation details |
 |
 |
spinal cord meninges |
 |
cerebral cortex ventricular layer |
 |
midbrain meninges |
|
 |
diencephalon meninges |
 |
telencephalon meninges |
 |
hindbrain meninges |
|
|
EMAGE:20037
view entry |
wild-type |
- |
ISH |
 |
|
|
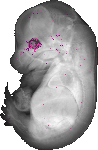
Ckmt1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.010 |
| Go to annotation details |
 |
 |
glossopharyngeal IX ganglion |
 |
vestibulocochlear VIII ganglion |
 |
vagus X ganglion |
|
 |
trigeminal V ganglion |
 |
nasal cavity olfactory epithelium |
 |
rectum |
 |
ventral grey horn |
|
|
EMAGE:10918
view entry |
wild-type |
- |
ISH |
 |
|
|
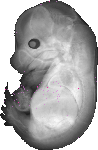
D930030D11Rik |
 |
 |
 |
TS23 |
14.5 dpc |
0.009 |
- |
EMAGE:16085
view entry |
wild-type |
- |
ISH |
 |
|
|
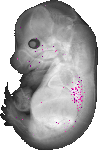
Lamc2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.009 |
| Go to annotation details |
 |
 |
lower jaw molar |
 |
upper jaw molar |
 |
thyroid gland |
|
 |
oral epithelium |
 |
naris |
 |
palatal shelf |
 |
submandibular gland primordium |
 |
medulla oblongata basal plate mantle layer |
 |
upper jaw incisor |
 |
esophagus |
 |
urethra of male |
 |
bladder |
 |
lower jaw incisor |
 |
metanephros |
 |
lung |
|
|
EMAGE:8993
view entry |
wild-type |
- |
ISH |
 |
|
|
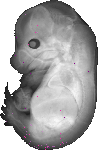
Dstyk |
 |
 |
 |
TS23 |
14.5 dpc |
0.009 |
|
EMAGE:8459
view entry |
wild-type |
- |
ISH |
 |
|
|
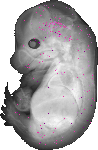
Mir511 |
 |
 |
 |
TS23 |
14.5 dpc |
0.009 |
|
EMAGE:16743
view entry |
wild-type |
- |
ISH |
 |
|
|
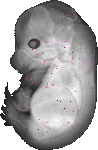
Slc35a5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.009 |
- |
EMAGE:6702
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file