|
|
Mrps11 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:19266
view entry |
wild-type |
- |
ISH |
 |
|
|
6430706D22Rik |
 |
 |
 |
TS23 |
14.5 dpc |
0.324 |
- |
EMAGE:19467
view entry |
wild-type |
- |
ISH |
 |
|
|
Hjurp |
 |
 |
 |
TS23 |
14.5 dpc |
0.324 |
- |
EMAGE:19468
view entry |
wild-type |
- |
ISH |
 |
|
|
Emg1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.323 |
- |
EMAGE:19368
view entry |
wild-type |
- |
ISH |
 |
|
|
Map2k3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.322 |
- |
EMAGE:9834
view entry |
wild-type |
- |
ISH |
 |
|
|
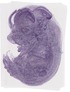
Snrpb2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.321 |
|
EMAGE:10965
view entry |
wild-type |
- |
ISH |
 |
|
|
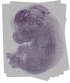
Hirip3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.320 |
- |
EMAGE:19704
view entry |
wild-type |
- |
ISH |
 |
|
|
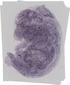
Lap3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.320 |
| Go to annotation details |
 |
 |
telencephalon ventricular layer |
 |
diencephalon lateral wall ventricular layer |
 |
olfactory cortex ventricular layer |
|
 |
hypothalamus ventricular layer |
 |
midbrain ventricular layer |
|
|
EMAGE:19379
view entry |
wild-type |
- |
ISH |
 |
|
|
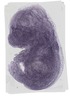
Ddx50 |
 |
 |
 |
TS23 |
14.5 dpc |
0.319 |
|
EMAGE:17543
view entry |
wild-type |
- |
ISH |
 |
|
|
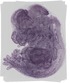
Nop2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.319 |
| Go to annotation details |
 |
 |
olfactory cortex ventricular layer |
 |
telencephalon ventricular layer |
 |
vibrissa |
|
 |
nasal cavity olfactory epithelium |
 |
upper jaw molar |
 |
thymus primordium |
 |
submandibular gland primordium |
 |
midbrain ventricular layer |
 |
vertebral axis musculature |
 |
upper jaw incisor |
 |
lower jaw molar |
 |
lower jaw incisor |
 |
liver lobe |
 |
renal cortex |
|
|
EMAGE:19479
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file